An experience that happened a few times in my career (not anymore, as I’ll explain) goes like this:
Collaborator: We have this data, can you tell us whether it fits our hypothesis?
Me: Sure
[ … data analysis happens … ]
Me: Nope, your data is no good. You need more/better data.
Collaborator: Oh, but such and such publication/former collaborator got different results.
Me: Can I see the reference?
Collaborator: Sure, here.
[ … reading happens … ]
Me: yeah, their results are bogus/cheated. Here’s why: [explanation]
Collaborator: I see a technical disagreement here, but I prefer the other guy’s results so we’re going to go with them…
This really doesn’t happen anymore because as a PI with a decent publication record, I now have enough status that the collaborators will believe me when I tell them that some other method’s results are bad.1
On the way to build the AMPSphere, we had to develop Macrel, which is a simple machine learning tool to classify peptides as despite the fact that this field already has >50 tools out there. They were all bad in the exact same way: too many false positives!
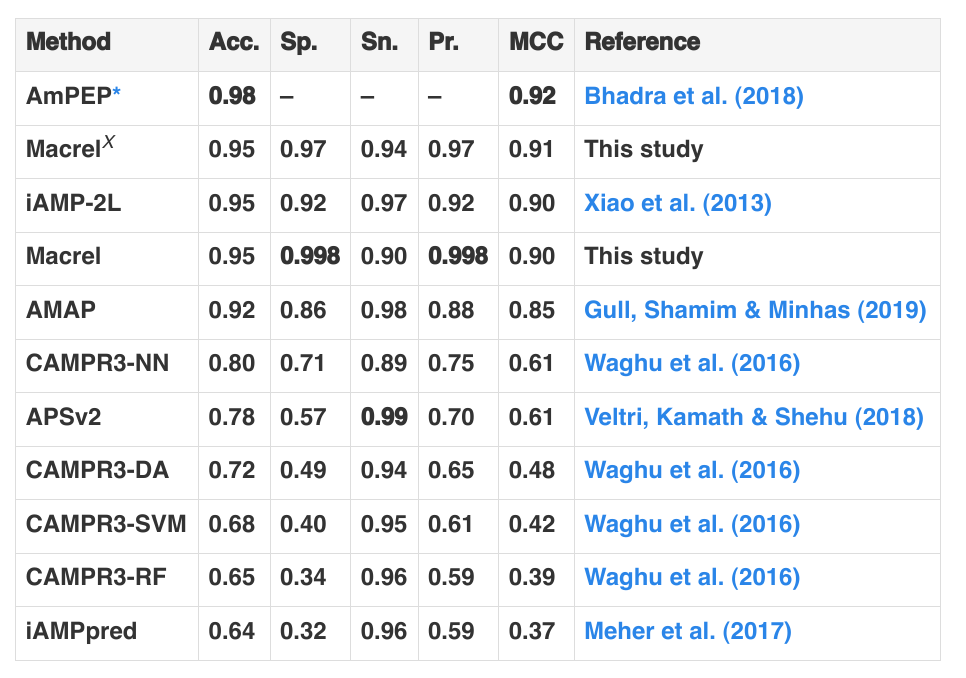
Here’s the results table from the Macrel manuscript:
You can see that other tools have higher Accuracy, but Macrel has the highest specificity (lowest false positive rate). But the accuracy is measured on a balanced test set, with equal numbers of negatives and positives. This matches almost no real-world use and, by standard, Bayesian reasoning means that if you apply the classifier in a more realistic setting (where there is a preponderance of negative compared to positive cases), most of the results will be false positives.
What is going wrong here? Why is there a vast literature of tools that are effectively unusable for their stated purpose. At the root of it all, the issue is one of incentives. We wanted a tool that would allow us to filter a huge collection of potential candidates (4.5 billion sequences) and get a smaller number that we would test in the lab. Therefore, high false positive rates would be a killer.
Too many users of these tools actually want something that allows them to weave a story for a paper. This is the true purpose of these tools.2 If you run an AMP prediction on your genome and it returns some hits, that seems better than a tool that finds nothing and, unless you are in the minority who is going to test these hits experimentally, you might not care very strongly if they are false positives. So, tool developers, particularly tool developers who must build careers as tool developers, need to cater to this bias in favor of false positives. We were building a tool with the goal of being users ourselves.
The result is that we fundamentally care less about having a lot of citations to Macrel3 than we care about Macrel being good! Ideally these would be the same, but I think they are not. I think that simply optimising for highest usage and short-term user satisfaction would have led us to output more false positives.

One of the few instances in my life where I said “please take me off this paper” was this exact situation. I told a collaborator that their data was not good enough (they needed to sequence deeper) and they got a different person to run a few tools that gave them a lot of false positives. They then wrote a manuscript claiming things that I think were false with this alternative analysis that still followed all the rules to get published: they used existing, seemingly well-validated, computational tool that, alas, returned a false positive result in the direction that made sense for that experiment.
The purpose of a system is what it does.